A Genetic Approach to the ApoE4 Puzzle
Quick Links
This paper was retracted on 17 June 2015.
For decades, scientists have wondered how ApoE4, the strongest genetic risk factor for Alzheimer’s disease, influences pathology. They know the allele affects cells and tissues in many ways, but which actually lead to AD? Researchers led by Asa Abeliovich at Columbia University, New York, outline a new approach to answer the question in the July 24 Nature online. They compared transcriptomes of healthy ApoE4 carriers to those of LOAD patients who do not carry this gene, and found common patterns of gene expression. Genes regulating endocytosis and the processing of the amyloid β precursor protein (APP) stood out, hinting that ApoE4 primes the brain for APP-related changes that lead to AD, suggested the authors. “The data jibe with longstanding human and mouse data showing that ApoE4 enhances brain amyloid pathology,” wrote Samuel Gandy, Mount Sinai Medical Center, New York to Alzforum in an email. “It is heartening to think that after 20 years of ApoE4 linkage to AD, there are new exciting leads that might offer an explanation.” However, he noted some caveats (see full comment below).
Studies of humans, mice, and cultured cells suggest that ApoE4 impairs Aβ degradation and clearance, and encourages inflammation and endocytosis, which create even more Aβ (see Castellano et al., 2011; Koistinaho et al., 2004; Zhu et al., 2012; and He et al., 2007). However, it is unclear if all or any of these effects cause AD, result from it, or even relate to the disease, said co-first author Herve Rhinn. He and colleagues wanted to figure out what changes occur first as a way of getting closer to the root causes of the disease.
To do that, Rhinn and co-first author Ryousuke Fujita looked for transcript alterations in the cerebral cortex of postmortem brains from 185 healthy people grouped by ApoE status. The analysis revealed more than 8,000 genes that were expressed differently in people who carried the ApoE4 allele compared to those with ApoE2 or E3. Of those hits, 215 turned up in a separate analysis comparing expression in 86 LOAD patients and 67 age-matched controls, all without ApoE4. These genes may be key to AD risk endowed by ApoE, suggested the authors.
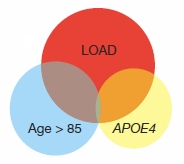
Genetic overlap among aging, ApoE4, and LOAD.
Image courtesy Herve Rhinn and Nature
Curiously, the ApoE4 pattern barely overlapped with that seen when the researchers compared gene expression among 64 older (age 85 and up) and 56 younger brains (age 75 or less). This suggested to the authors that ApoE and aging—the major risk factor for late onset AD (LOAD)—contribute to the disease via separate mechanisms (see figure below). The implication is that “different genetic forms or subtypes of Alzheimer’s disease may have specific underlying mechanisms and respond differently to treatment,” wrote Vivek Swarup and Daniel Geschwind of the University of California, Los Angeles, in an accompanying editorial.
To find out what mediated the differences in expression, the researchers applied a systems biology approach called differential co-expression analysis (DCA), which weighs results based on co-regulated genes (see ARF related news story). This identified 20 "master regulators" of the 215 genes. Researchers had previously associated the top hit, the zinc-binding protein RNF219, with cognitive performance, lipid metabolism, and brain ventricular volume. Another candidate—the synaptic vesicle protein SV2A—helps regulate neuronal endocytosis. Scientists had previously tied several of the regulators, including APBA2, ITM2B, and FYN, to the processing and intracellular trafficking of APP. Fyn kinase has also been linked to Aβ toxicity via tau (see ARF related news story).
To test whether these 20 genes bring about ApoE4’s effects, Rhinn and colleagues used small hairpin RNAs to knock down several individually in mouse N2a neuroblastoma cells and gauge how the cells reacted to added human recombinant ApoE4. This treatment spurred production of Aβ40 and Aβ42 in wild-type N2a cells, but not when the genes were knocked down.
The researchers then took a closer look at RNF219 and SV2A. Knocking down either prevented ApoE4 from boosting β–secretase 1 (BACE1) processing of APP, which it does in wild-type cells by promoting co-localization of BACE1 and APP in endosomes. Levetiracetam, an SV2A inhibitor used to treat epileptic seizures, also reduced Aβ production in N2a cells as well as in neurons induced from human fibroblasts donated by ApoE4 carriers. Together, these results suggest that RNF219 and SV2A mediate ApoE4’s effects on APP processing, concluded the authors.
Previous studies suggested that levetiracetam lowers neuronal hyperactivity and improves cognition in J20 mice. Interestingly, other anti-epileptic drugs, which have no affinity for SV2A, failed to achieve the same result (see ARF related news story). “We think levetiracetam works by reducing Aβ,” Abeliovich told Alzforum. “Our work reframes existing data, and suggests that hitting this target [SV2A] could be disease-altering,” he said. Michela Gallagher, Johns Hopkins University, Baltimore, Maryland, who previously reported that levetiracetam improved memory in people with mild cognitive impairment (see ARF related news story), agreed. “This report has potential therapeutic implications in support of SV2A as a target to prevent LOAD, at least in ApoE4 carriers,” Gallagher wrote to Alzforum in an email (see full comment below). She pointed out that benefits of levetiracetam extend to mice and humans that do not carry ApoE4, so the drug could help a broader population.
Scott Small, also at Columbia University, pointed out that the data support a hypothesis, summarized in his review written with Gandy, that any defect that keeps APP and BACE1 in endosomes will increase Aβ production and drive pathology (see Small and Gandy, 2006).
Other experts cautioned that the paper was still speculative, and required more exploration. “It will be interesting to see if the function of SV2A and RNF219 described here in cell culture can be reproduced in AD animal models,” wrote Lars Ittner, University of New South Wales, Sydney, Australia, to Alzforum via email (see full comment below). Ittner pointed out that how these ApoE4-driven changes relate to tau remains to be seen. Gandy cautioned that in other cultured neural cells, ApoE4 does not modify APP sorting or processing (see Biere et al., 1995).
Abeliovich plans to test if and how the other hits from this DCA study influence disease progression. In addition, he will extend this analysis beyond ApoE4 to other risk variants identified in genome-wide association studies for AD, and apply the method to Parkinson’s disease (see ARF related news story).—Gwyneth Dickey Zakaib
References
News Citations
- ApoE4 Paper Retracted from Nature
- A Genetic Approach to the ApoE4 Puzzle
- Bioinformatics Uncovers Pathological Synuclein RNA
- Honolulu: The Missing Link? Tau Mediates Aβ Toxicity at Synapse
- Anticonvulsants Reverse AD-like Symptoms in Transgenic Mice
- Epilepsy Drug Calms the Hippocampus, Aids Memory
Paper Citations
- Castellano JM, Kim J, Stewart FR, Jiang H, DeMattos RB, Patterson BW, Fagan AM, Morris JC, Mawuenyega KG, Cruchaga C, Goate AM, Bales KR, Paul SM, Bateman RJ, Holtzman DM. Human apoE isoforms differentially regulate brain amyloid-β peptide clearance. Sci Transl Med. 2011 Jun 29;3(89):89ra57. PubMed.
- Koistinaho M, Lin S, Wu X, Esterman M, Koger D, Hanson J, Higgs R, Liu F, Malkani S, Bales KR, Paul SM. Apolipoprotein E promotes astrocyte colocalization and degradation of deposited amyloid-beta peptides. Nat Med. 2004 Jul;10(7):719-26. PubMed.
- Zhu Y, Nwabuisi-Heath E, Dumanis SB, Tai LM, Yu C, Rebeck GW, LaDu MJ. APOE genotype alters glial activation and loss of synaptic markers in mice. Glia. 2012 Apr;60(4):559-69. PubMed.
- He X, Cooley K, Chung CH, Dashti N, Tang J. Apolipoprotein receptor 2 and X11 alpha/beta mediate apolipoprotein E-induced endocytosis of amyloid-beta precursor protein and beta-secretase, leading to amyloid-beta production. J Neurosci. 2007 Apr 11;27(15):4052-60. PubMed.
- Small SA, Gandy S. Sorting through the cell biology of Alzheimer's disease: intracellular pathways to pathogenesis. Neuron. 2006 Oct 5;52(1):15-31. PubMed.
- Biere AL, Ostaszewski B, Zhao H, Gillespie S, Younkin SG, Selkoe DJ. Co-expression of beta-amyloid precursor protein (betaAPP) and apolipoprotein E in cell culture: analysis of betaAPP processing. Neurobiol Dis. 1995 Jun;2(3):177-87. PubMed.
Further Reading
No Available Further Reading
Primary Papers
- Swarup V, Geschwind DH. Alzheimer's disease: From big data to mechanism. Nature. 2013 Aug 1;500(7460):34-5. PubMed.
- Rhinn H, Fujita R, Qiang L, Cheng R, Lee JH, Abeliovich A. Integrative genomics identifies APOE ε4 effectors in Alzheimer's disease. Nature. 2013 Aug 1;500(7460):45-50. PubMed. RETRACTED
Annotate
To make an annotation you must Login or Register.

Comments
Johns Hopkins Universiy
Apart from aging itself, apolipoprotein E (Apo-E) polymorphic alleles are the primary genetic determinants of risk for sporadic late onset Alzheimer disease (LOAD). Rhinn et al. have uncovered a candidate effector pathway for Apo-E4 on amyloid beta precursor protein (APP) endocytosis and metabolism contributing to LOAD. In addition to advancing biological understanding, the report has potential therapeutic implications in support of Sv2a as a target to prevent LOAD, at least in ApoE4 carriers. Would the Sv2a inhibitor used to suppress altered APP processing in APOE4-positive human induced neurons (hiNs) have in vivo efficacy in that population?
While that remains to be seen, it is important to note that targeting Sv2a has shown beneficial effects in aging, MCI patients, and AD models apart from ApoE4. This is true for our (Bakker et al., 2012) and a second prior study using levetiracetam cited by the authors (Sanchez et al., 2012).
The AD mouse model used by Sanchez et al. is devoid of ApoE4 but showed improved cellular, network, and behavioral outcomes specific for mice with hAPP overexpression after treatment with the Sv2a ligand levetiracetam (see also Spiegel et al., 2013 for further evidence of a therapeutic benefit in a rodent model, absent ApoE4). Hyperactivity leading to network dysfunction is a common feature of these preclinical models. As Rhinn and colleagues noted, our report using low dose levetiracetam in amnestic mild cognitive impairment (MCI) ascribed therapeutic benefit to effective reduction of hippocampal hyperactivity. We have since completed additional cohorts of MCI patients under the same protocol (totaling N=54). In the aggregate MCI study population, 43 percent of the genotyped subjects were APOE4 carriers. The signature of hyperactivity localized within the hippocampal formation was not restricted to APOE4 carriers but was evident in carriers and non-carriers alike. The notion that this condition contributes to disease progression is supported by a close association between the magnitude of hippocampal hyperactivity in MCI with the severity of structural atrophy in key AD related areas of the brain (Putcha et al., 2011). It is also notable that hippocampal hyperactivity is seen pre-symptomatically in early onset AD (Quiroz et al., 2010) as well as in asymptomatic APOE4 carriers and in aMCI, irrespective of E4 carrier status. Hence, drugs that target Sv2a could be beneficial therapeutically in the broader context of prodromal AD, while not being limited to the ApoE4 population.
References:
Bakker A, Krauss GL, Albert MS, Speck CL, Jones LR, Stark CE, Yassa MA, Bassett SS, Shelton AL, Gallagher M. Reduction of hippocampal hyperactivity improves cognition in amnestic mild cognitive impairment. Neuron. 2012 May 10;74(3):467-74. PubMed.
Sanchez PE, Zhu L, Verret L, Vossel KA, Orr AG, Cirrito JR, Devidze N, Ho K, Yu GQ, Palop JJ, Mucke L. Levetiracetam suppresses neuronal network dysfunction and reverses synaptic and cognitive deficits in an Alzheimer's disease model. Proc Natl Acad Sci U S A. 2012 Oct 16;109(42):E2895-903. PubMed.
Spiegel AM, Koh MT, Vogt NM, Rapp PR, Gallagher M. Hilar interneuron vulnerability distinguishes aged rats with memory impairment. J Comp Neurol. 2013 Jun 8; PubMed.
Putcha D, Brickhouse M, O'keefe K, Sullivan C, Rentz D, Marshall G, Dickerson B, Sperling R. Hippocampal hyperactivation associated with cortical thinning in Alzheimer's disease signature regions in non-demented elderly adults. J Neurosci. 2011 Nov 30;31(48):17680-8. PubMed.
Quiroz YT, Budson AE, Celone K, Ruiz A, Newmark R, Castrillón G, Lopera F, Stern CE. Hippocampal hyperactivation in presymptomatic familial Alzheimer's disease. Ann Neurol. 2010 Dec;68(6):865-75. PubMed.
View all comments by Arnold BakkerIcahn School of Medicine at Mount Sinai
The data jibe well with longstanding human and mouse data showing that APOE4 enhances brain amyloid pathology robustly (especially cerebral amyloid angiopathy). It is heartening to think that after 20 years of APOE4 linkage to AD, there are new exciting leads that might, at long last, offer an explanation, especially since APOE4 is present in about 50 percent of patients with AD and is the most common identified genetic risk for the disease. The claim that APOE isotype-specific modulation of the expression of protein sorting genes can be demonstrated using patient-specific iPS cell technology certainly comports well with the generally agreed concept that APP sorting—specifically endosomal residence time—plays a critical role in amyloid β generation.
I have some serious concerns about this paper, however:
First, overexpression of APOE4 vs APOE3 cDNAs in cultured cells does not typically change APP sorting or Aβ biogenesis. This has been tested before by Selkoe and Younkin (see Biere et al., 1995) and by others, including our lab.
Second, some of the genes that the authors link to APOE4 have been linked to AD and have been shown to regulate APP processing and Aβ biogenesis but those actions have been associated with either missense mutations or polymorphisms that appear to act by modulating gene expression. APOE4 was not required to reveal the effects of the mutations/polymorphisms on APP processing and Aβ biogenesis in cells nor has APOE4 been required for the mutations/linkages to exert effects on disease risk.
The conventional wisdom on the likely mechanism of APOE4’s effects on AD pathogenesis favors APOE-isotype-specific modulation of Aβ clearance. Randy Bateman’s in vivo human data support this model (see ARF related news story). One conceivable (albeit unlikely) scenario that would unify the existing data is that the APOE4-misregulated genes that the authors report are not only APP missorting genes but also function in the uptake and clearance of secreted Aβ. This possibility remains to be tested.
Third and lastly, the use of recombinant ApoE also raises a red flag. Only astrocytes express and lipidate ApoE and package it into discs and particles in a brain-relevant physiological fashion. Biovision recombinant ApoE is produced in bacteria and lyophilized. Apolipoproteins are extraordinarily difficult reagents to generate and handle properly. Many cell biologists would say that the only physiologically relevant ApoE currently employed in brain and AD research is that recovered from the conditioned media of David Holtzman's immortalized APOE-isotype-expressing astrocytes. Bacterially expressed, unlipidated, lyophilized ApoE is about as far from physiologically relevant as the chloroform: methanol extracted and purified material that has been responsible for two decades worth of artifacts in the literature.
References:
Biere AL, Ostaszewski B, Zhao H, Gillespie S, Younkin SG, Selkoe DJ. Co-expression of beta-amyloid precursor protein (betaAPP) and apolipoprotein E in cell culture: analysis of betaAPP processing. Neurobiol Dis. 1995 Jun;2(3):177-87. PubMed.
View all comments by Sam GandyJohns Hopkins University
I read this paper with keen interest yesterday, but also found myself uneasy with the recombinant ApoE cell treatments as noted by Sam Gandy. Following my reading of his comments today, I took another look at that paper with a different view, asking if these points diminished my initial enthusiasm. I arrived at the conclusion that even if all of the recombinant ApoE treatment studies were removed from the paper as potentially artifactual, the conclusions of the authors still stand on the basis of the differential co-expression analyses and the experiments with the human induced neuronal cultures.
University of New South Wales
Rhinn and colleagues report in this very interesting paper similar differential gene regulation in disease-free APOE4 carriers and non-APOE4 late onset Alzheimer’s disease (AD) patients, using differential co-expression analysis. The overlap in gene expression identified a number of modifiers of APP processing and transport, including SV2A and RNF219, which has not been linked to AD before. The authors move on to show in cell culture that both SV2A and RNF219 regulate the release of Aβ from human APP- expressing cells in an APOE4-dependent manner. Amongst the identified genes was also FYN, which we linked to Aβ toxicity and tau in our previous work (Ittner et al., 2010).
However, whether the modulation of Aβ pathology in the context of APOE4 also involves tau remains to be seen. Interestingly, the gene profile analysis suggests there are independent mechanisms at play in APOE4-positive late-onset AD and age-related late-onset AD. It will be interesting to see if the function of SV2A and RNF219 described here in cell culture can be reproduced in AD animal models, though there is already supporting evidence from AD mouse studies for SV2A, a target of levetriacetam.
References:
Ittner LM, Ke YD, Delerue F, Bi M, Gladbach A, van Eersel J, Wölfing H, Chieng BC, Christie MJ, Napier IA, Eckert A, Staufenbiel M, Hardeman E, Götz J. Dendritic function of tau mediates amyloid-beta toxicity in Alzheimer's disease mouse models. Cell. 2010 Aug 6;142(3):387-97. Epub 2010 Jul 22 PubMed.
View all comments by Lars M. IttnerMake a Comment
To make a comment you must login or register.