γ-Secretase: Ins and Outs of a Voracious Membrane Protein Grinder
Quick Links
Several researchers reported their latest data on the γ-secretase complex and APP processing. This research has expanded greatly beyond an initially narrow focus on Aβ production to where it now represents a lively subarea of cell biology that explores how aspartyl proteases carrying a GxGD motif carry out regulated intramembraneous proteolysis (RIP) of type 1, or single-pass, membrane proteins. γ-secretase remains the most famous representative of this group of proteases, but the related, and somewhat simpler, family of signal peptide peptidases (SPPs) is becoming better known and offers useful comparisons for the study of γ-secretase. New trends in this area include a growing realization that γ-secretases and SPPs are using a similar catalytic mechanism, whereby multiple cleavages inside the membrane give rise to a multitude of different fragments, Aβ40 and 42 being but two of them. This mechanism is not processive like that of a peptidase, which chops down its substrate from one end; rather, the substrate moves through the membrane stepwise and gets cleaved in particular positions.
Researchers are beginning to view γ-secretase as a membrane proteasome, or even as a sort of kitchen sink garbage disposal, whose job it is to remove residual transmembrane stubs of many different membrane receptors and proteins, such as APP. In parallel to that activity, though, γ-secretase also carries out highly specific cuts necessary for nuclear signaling. As the list of γ-secretase substrates continues to grow past the 25 or so known to date, researchers are realizing that this protease cares less about the amino acid sequence of its substrates than typical proteases do. Rather, it primarily requires that the previous step of α or β cleavage, also called ectodomain shedding, generate a fragment of a particular length that sports a free N-terminal. Beyond that, the question of how substrate specificity is conferred remains a puzzle.
Another big question concerns regulation. To some investigators, γ-secretase and SPPs look to be rather indiscriminate beasts that cut constitutively once suitable substrates are present. To them, most of the finely tuned regulation and important biology appears to occur upstream, at the level of α cleavage. That said, a brand new secretase complex member and ongoing work on lipids suggest that there is much more to γ-secretase modulation than presently meets the eye.
Finally, one emerging theme concerns comparisons with channel proteins. First glimpses of the complex’s putative 3D structure, and cell biology studies of the way it assembles, are reminding scientists of known aspects of ion channels or the sec61 protein pore through which type 1 membrane proteins slide on their way off the ribosome and into the ER. (Truth be told, this notion has been around for a while; see, e.g., Fred Van Leuven’s comment, and the idea is now gaining new data.) The Eibsee conference offered news on all these fronts. Below find presentation summaries.
Gang Yu, of the University of Texas Southwestern Medical Center in Dallas, started things off by recapping his recent paper showing that γ-secretase action begins when the complex member nicastrin recognizes the substrate (see news and discussion at Shah et al., 2005). Yu found that α or β cleavage of a given substrate protein, for example, APP or Notch, will expose a membrane-bound stub of usually less than 100 amino acids with a new free N-terminus. (In the confusing terminology of RIP, these stubs are called C99/C100/βCTF for APP, NEXT for Notch.) It is this stub that nicastrin latches onto and then swivels around, a bit like a construction crane, to move the substrate deeper into the γ-secretase complex for docking and cutting. Nicastrin forwards the substrate in this way by way of its ectodomain, whereas its transmembrane domain plays a role in the assembly and the activity of the γ-secretase complex. (see image below). Importantly, this crane-like ectodomain of nicastrin appears to select potential substrates more by their length and the presence of the free N-terminal stub than by their sequence, Yu said. Nicastrin uses a conserved DAP domain and, in particular, a glutamate at amino acid position 333 to interact with the substrate, and this site on nicastrin is indeed required for γ-secretase activity. In studying nicastrin’s modus operandi, Yu noted some similarities to how amino peptidases function. These are enzymes that catalyze the hydrolysis of the terminal peptide bond at the amino end of a polypeptide. This study proposes a common mechanism for how nicastrin recognizes all γ-secretase substrates, but it remains to be experimentally tested with regard to most substrates besides APP and Notch. The reality will likely prove to be more complex than this current model, Yu noted.
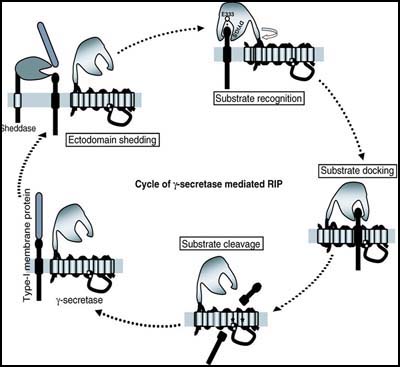
Nicastrin, via its ectodomain, recognizes the new free N-terminus of the substrates (and via its TM domain, participates in γ-secretase assembly and activity) [Figure courtesy of Gang Yu]
Harald Steiner, of Ludwig Maximilians University in Munich, first recapped some milestones in γ-secretase research, including its reconstitution in yeast (Edbauer et al., 2003) and, since then, in various other cell types. Later, it turned out that distinct γ-secretase complexes exist, made up by different combinations of the presenilin and Aph isoforms plus nicastrin and Pen-2 (Shirotani et al, 2004; Serneels et al., 2005). A pressing goal at this point is to find out exactly how the enzyme complex distinguishes APP from Notch (and other substrates) so that this information can help guide the search for new γ-secretase modulator drugs that can tweak APP cleavage selectively. At the Eibsee conference, talks and informal discussion suggested that several competing groups appear to be converging on the same spot on the protease. In his keynote lecture, Selkoe recounted how his group’s pursuit of this site developed from follow-up studies of Paul Greengard’s discovery that the cancer drug Gleevec inhibits cleavage of C100 (the APP substrate generated by BACE cleavage) but not of the corresponding Notch substrate (Netzer et al., 2003). Certain kinase inhibitors turned out to do the same thing, as does ATP, and these compounds all appear to act on a nucleotide-binding site on γ-secretase (Fraering et al., 2005). Michael Wolfe, at Brigham Hospital, has proposed a site for substrate docking that lies near the active site. It might serve as a stopover between the substrate’s initial binding to nicastrin at the surface of the protease and its eventual cleavage deep inside the protease (Kornilova et al., 2005).
For his part, Steiner is zeroing in on the same site. Coming originally from an interest in how the substrate transits from nicastrin to the active site, Steiner, Haass, and colleagues are finding that substrate docking and distinguishing between APP and Notch may occur in that very same place near the active site. When the scientists took an evolutionarily distant, roundworm cousin of γ-secretase and swapped its active site with that of presenilin-1, the chimera processed APP just fine but could not cut Notch. In-vivo tests in a C. elegans Notch mutant confirmed that something about this presenilin-1 active site was specific to APP. Mutation in its transmembrane domain 7 revealed this something to be a single amino acid, namely a leucine at position 383. This residue represents the X position of a GXGD motif on this domain that lies next to the catalytic site. Steiner’s work suggests that the final substrate selection happens there, as Wolfe had proposed. In effect, the GXGD motif might be a final “gate” to the active site, Steiner said (Yamasaki et al., 2006). The motif occurs in γ-secretase and its relatives, the signal peptide peptidases, and its “D” actually represents one of the two proposed catalytic aspartates, implying that GXGD encompasses both substrate selection and part of the active site. Its role, if any, in the selection of other γ-secretase substrates is not clear yet.
In the discussion, Selkoe speculated that a Walker-like motif exists on transmembrane domain 7 of presenilin that might account for ATP binding and represent a site of allosteric regulation of γ-secretase. Described by John E. Walker as sequence motifs indicating the presence of nucleotide binding domains, these sequences typically occur in protein regions accessible to water so they can support ATP hydrolysis. It is unusual for such a motif to lie hidden inside a hydrophobic region of an intramembranous enzyme complex, but then, most everything about γ-secretase is unusual. The presence of such a motif might yield clues to γ-secretase function, that is, where water enters into the lipid bilayer to enable breaking of the substrate’s peptide bonds, and how the catalytic aspartates in presenilin’s transmembrane domains become protonated in preparation for proteolysis, Selkoe noted.
This Walker-like motif showed up in yet another guise. By studying APP dimerization, Gerd Multhaup arrived at this region, as well. With postdoc Lisa Munter, Multhaup discovered that it turns out to harbor a third, potentially critical dimerization spot on APP that influences the sequential cleavages of the γ-secretase substrate. By doing so, it helps determine what forms of Aβ come out of this process.
APP dimerization itself, by two prior points of contact between monomers, had been published before (see, for example, Beher et al., 1996; Scheuermann et al., 2001). Some other γ-secretase substrates are thought to dimerize, as well. At the conference, Multhaup started out by saying that his group had discovered a third, new site of contact between two APP molecules (as well as other substrates) that mediates dimerization. A FRET study confirmed dimerization in living cells. On APP, this new site includes three consecutive GxxxG motifs. From other research, such motifs are known to promote interaction of protein helices, and a recent Alzforum Discussion challenged the field to ponder whether they somehow might mediate Aβ toxicity. On APP, one of the GxxxG motifs within the Aβ sequence represents a particularly important site for APP dimerization and lies within the Walker-like motif. Multhaup speculated further that the generation of Aβ42 depends on the GxxxG motifs by influencing APP-APP interaction. Further details on this work will be reported upon publication.—Gabrielle Strobel.
References
News Citations
Webinar Citations
Paper Citations
- Shah S, Lee SF, Tabuchi K, Hao YH, Yu C, LaPlant Q, Ball H, Dann CE, Südhof T, Yu G. Nicastrin functions as a gamma-secretase-substrate receptor. Cell. 2005 Aug 12;122(3):435-47. PubMed.
- Edbauer D, Winkler E, Regula JT, Pesold B, Steiner H, Haass C. Reconstitution of gamma-secretase activity. Nat Cell Biol. 2003 May;5(5):486-8. PubMed.
- Shirotani K, Edbauer D, Prokop S, Haass C, Steiner H. Identification of distinct gamma-secretase complexes with different APH-1 variants. J Biol Chem. 2004 Oct 1;279(40):41340-5. PubMed.
- Serneels L, Dejaegere T, Craessaerts K, Horré K, Jorissen E, Tousseyn T, Hébert S, Coolen M, Martens G, Zwijsen A, Annaert W, Hartmann D, De Strooper B. Differential contribution of the three Aph1 genes to gamma-secretase activity in vivo. Proc Natl Acad Sci U S A. 2005 Feb 1;102(5):1719-24. PubMed.
- Netzer WJ, Dou F, Cai D, Veach D, Jean S, Li Y, Bornmann WG, Clarkson B, Xu H, Greengard P. Gleevec inhibits beta-amyloid production but not Notch cleavage. Proc Natl Acad Sci U S A. 2003 Oct 14;100(21):12444-9. PubMed.
- Fraering PC, Ye W, LaVoie MJ, Ostaszewski BL, Selkoe DJ, Wolfe MS. gamma-Secretase substrate selectivity can be modulated directly via interaction with a nucleotide-binding site. J Biol Chem. 2005 Dec 23;280(51):41987-96. PubMed.
- Kornilova AY, Bihel F, Das C, Wolfe MS. The initial substrate-binding site of gamma-secretase is located on presenilin near the active site. Proc Natl Acad Sci U S A. 2005 Mar 1;102(9):3230-5. PubMed.
- Yamasaki A, Eimer S, Okochi M, Smialowska A, Kaether C, Baumeister R, Haass C, Steiner H. The GxGD motif of presenilin contributes to catalytic function and substrate identification of gamma-secretase. J Neurosci. 2006 Apr 5;26(14):3821-8. PubMed.
- Beher D, Hesse L, Masters CL, Multhaup G. Regulation of amyloid protein precursor (APP) binding to collagen and mapping of the binding sites on APP and collagen type I. J Biol Chem. 1996 Jan 19;271(3):1613-20. PubMed.
- Scheuermann S, Hambsch B, Hesse L, Stumm J, Schmidt C, Beher D, Bayer TA, Beyreuther K, Multhaup G. Homodimerization of amyloid precursor protein and its implication in the amyloidogenic pathway of Alzheimer's disease. J Biol Chem. 2001 Sep 7;276(36):33923-9. PubMed.
Further Reading
Annotate
To make an annotation you must Login or Register.

Comments
No Available Comments
Make a Comment
To make a comment you must login or register.